Date Range:
Publication Date:
Owner/Creator:
Project:
Data sources:
Methods:
EXPERIMENTAL DESIGN AND METHODS: Samples are collected and processed by PIE researchers. For 2003-2006, sampling stations along the Plum Island Sound-Parker River were at fixed geographic locations at specific "Bends" in the river, reflected in the site name (top map). In 2008, we began sampling the water column in salinity space rather than at specific geographic locations along the river. This sampling approach was adopted in order to follow particular water masses in this macrotidal estuary. In practical terms, it means that sampling locations, or stations, are not static.
Samples are collected and processed by PIE researchers. For 2003-2006, sampling stations along the Plum Island Sound-Parker River were at fixed geographic locations at specific "Bends" in the river, reflected in the site name (top map). In 2008, we began sampling the water column in salinity space rather than at specific geographic locations along the river. This sampling approach was adopted in order to follow particular water masses in this macrotidal estuary. In practical terms, it means that sampling locations, or stations, are not static.
Therefore, we created four bounding boxes to encompass three regions of the river plus the sound (bottom map), and mapped the 11 sampling locations (latitude and longitude are logged at each station) from each transect along the mainstem of the estuary, so each station may be located within one of these bounding boxes as well as in salinity space. When needed, station coordinates have also been used to map the specific location of a station from a specific transect.
Bounding boxes: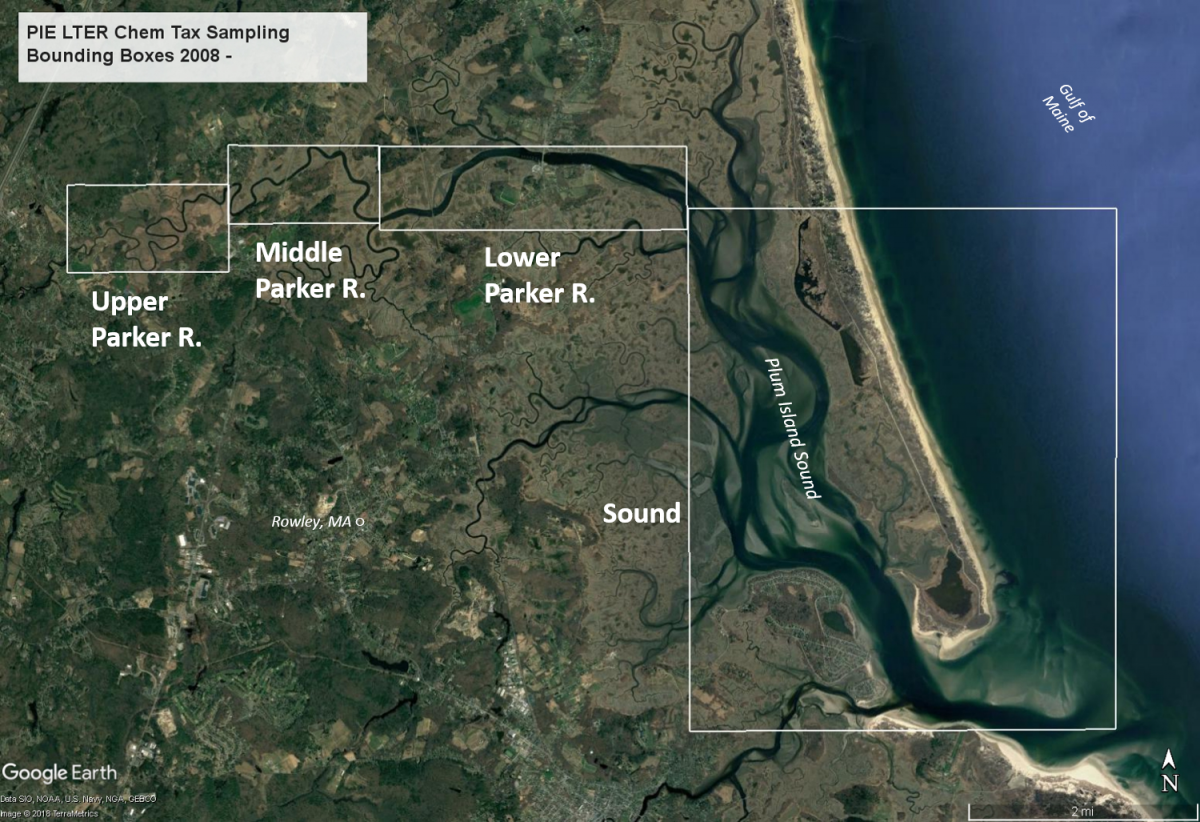
Sound (Plum Island Sound; EST-PR-SoundBND), 0 to ~9.5 km, proceeding upstream from the mouth of the sound; Lower Parker River (EST-PR-LowerParkerBND) , ~9.5 - 14.5 km; Middle Parker River (EST-PRMiddleParkerBND), ~14.5 - 18.75 km; Upper Parker River (EST-PR-UpperParker BND)., ~18.75 to 24.25 km (the Parker R. Dam).
HPLC and Chem Tax analyses are performed by the HPLC Photopigment Analysis Facility at the University of South Carolina, under the direction of Dr. James Pinckney. The procedure is similar to the standard method for filtering water samples for Chl a analysis by fluorometry or spectrophotometry. For HPLC analysis, the approach is to collect as much material as possible to provide high resolution of all the algal groups present and facilitate the confirmation of pigment identities by comparisons with absorption spectra for authentic standards.
The basic filtration steps are as follows:
1. Collect water (2 gallons for IBYC, Nelson, Station 24, Station 22 and 1 gallon for Stations 20, 18, 16, 12, 8, 4, 0) in a clean container. Keep the water cool and in the dark during transport and short-term storage (an insulated cooler with a little ice works well). The time from collection to filtration should be less than 3 hours. There will be a whole water sample filtered as described below, need 2 replicates or 2 filters per station. There will also be a < 20 um sample which is whole water prescreened through 20 um nitex where the filtrate will then be filtered as described below, need 2 replicates or 2 filters per station.
2. Using square-ended forceps (to prevent poking a hole in the filter), place a single Whatman GF/F filter (2.5cm) onto a Gelman filter funnel. Make sure the funnel forms a tight seal with the base.
3. The filter funnel should be attached to a vacuum system. We use a home-made PVC manifold that holds several funnels, and each funnel has a valve to open/close the vacuum. The manifold is attached to heavy-walled tygon tubing and a large vacuum flask (5 gallon glass water bottle). A second hose runs from the carboy to the vacuum pump. Use a moderate vacuum for the filtrations (ca. 100 to 200 mm Hg, or <7 in. Hg vac).
4. Open the vacuum and pour in a pre-measured amount (using a clean graduated cylinder) of sample water. This is the tricky part. You may have to use trial and error to determine how much water you can filter before the filter clogs. Samples closer to the mouth of Plum Island Sound will require larger volumes (500-1000ml) than samples up the Parker River (150-700ml) depending upon season and discharge. Filter water samples in duplicate, then pool the two filters for HPLC analyses. Be sure to record the total volume of water filtered for each duplicate sample.
5. Using square-tipped forceps, gently fold the filter in half, with the side containing the sample on the inside of the fold. Remove the filter, keeping it folded, and place the filter between two sheets of tissue paper (we use paper towels). Gently press the filter with your thumb to remove excess water from the filter. (Excess water in the filter reduces the extraction efficiency of the acetone solvent).
6. Label the outside of a small piece of aluminum foil (a square piece with dimensions of 5 x 5 cm) with the sample identification number. Use a black sharpie pen and make sure the ink dries (sharpie ink will destroy the pigment analysis….and the filters are magnets for sharpie ink!)
7. Place the duplicate filters together (keeping them folded in half) onto the foil and fold the foil so that the label appears on the outside. Make sure the filter is completely covered by the foil wrapper. Bend the edges of the foil to make sure the filter is sealed within the foil. Place the foil in a freezer (the colder the better). Transport to MBL in small ChemTax cooler box with ice pack. Keep the sample completely frozen and in the dark until you are ready to conduct the HPLC analysis. Samples can be stored for as long as 1 year at -80 C.
8. Fill out ChemTax Transect data sheet with information about samples.
9. For shipping t othe USC lab, the samples should be sent Express Overnight in a styrofoam cooler filled with dry ice. Fill any empty space with newspaper. Please let the lab know when you are shipping so that thety can be on the lookout for the shipment. Plan shipments to avoid weekend delays.
NOTES and COMMENTS
The initial pigment ratios used by ChemTax were based on published values for estuarine populations (Roy et al. 2011; Schlüter et al., 2000) selected to fit results of microscopy of a subset of duplicate samples (David Borkman, URI) from the Parker River estuary. Microscopy was important in both defining the groups and for qualitative evaluation of ChemTax. At PIE, the chlorophyll a apportioned to the combined category "Diatoms & Chrysophytes" is largely diatom chlorophyll. We are continuing the comparisons between CHEMTAX and microscopy, which may lead to further refinements to the calibration.
When samples for 2018 were submitted for pigment and CHEMTAX analysis, Dr. Pinckney judged it was appropriate to reanalyze the data in terms of CHEMTAX for all years, using the microscopy results and current taxonomic classifications to inform the analysis. This led to two changes in phytoplankton group categories (Variable Name): DiatomsandChrysophytes became Diatoms, and Prymesiopytes were omitted.
Taxa identified by microscopy (2008-2009):
Bacillariophyceae:
Diatoms, centric: Chaetoceros debilis , Chaetoceros spp. <10 um dia, Chaetoceros spp. 10-30 um dia, Chaetoceros spp. 10-30 um dia , Coscinodiscus spp. (30 - 60 um diameter) , Dactyliosolen
fragilissima , Odontella spp. , Skeletonema spp. , Stephanodiscus spp. , Thalassiosira rotula , Thalassiosira spp. <10 um dia, Thalassiosira spp. 10-30 um dia, unidentified Centric diatom <10 um dia
Diatoms, pennate: Amphora spp. ,Cocconeis scutellum , Cylindrotheca closterium , Gyrosigma spp., Licmophora abreviata, naviculoid diatom <10um, Pennate diatom <10 um L, Pennate diatom 10-30 um L,
Pennate diatom 30-60 um L , Pleurosigma aestuarii , Pleurosigma spp. , Pseudo-nitzschia pseudodelicatissima , Pseudo-nitzschia spp., Surirella spp., Synedra spp., Thalassionema nitzschioides
Dinophyceae: Ceratium lineatum , Gymnodinium spp. <20 um long , Gymnodinium spp. 20 - 40 um long, Heterocapsa rotundata , Heterocapsa triquetra, Prorocentrum minimum, Gyrodinium spp (20-40 um L),
Protoperidinium spp. (30-60 um diameter) , Protoperidinium spp. (30-60 um diameter), Scrippsiella trochoidea
Prasinophyceae: Pyramimonas spp. (Prasino A; could be clone PRD18DOI, AY948021; Prasino B may be reassigned to Chlorophytes, pending further refinement of the pigment matrix)
Chrysophyceae: Dinobryon spp., Calycomonas wulfii (both ChrysoA; ChrysoB types have not been identified at PIE, but an example would be Pelagococcus subviridi)
Euglenophyceae: Eutreptia / Eutreptiella spp.
Cryptophyceaes: Cryptomonads <20 um L
microflagellates: unidentified phytoflagellates<10 um diameter (likely includes Prasinophytes identified by CHEMTAX)
Dictychophyceae (silicoflagellate): Dictyocha speculum (has pigment composition similar to HaptophyteB)
Raphidophyceae: not yet identified at PIE; but could include Heterosigma, Fibrocapsa
Cyanobacteria: unidentified Nostoc-like species.
Prymnesiophyceae: Phaeocysitis (Hapto-B) (Phylum Haptophyta)
Maintenance:
On going collections
Version 01: 17Sep2013, metadata and data created to comply with importation to Drupal and LTER PASTA. Used MarcrosExportEML_HTML (working)pie_excel2007_Sep2013.xlsm 9/10/13 12:02 PM for QA/QC to EML 2.1.0
Version 02: March 8, 2016, data and metadata updates to comply with importation to Drupal and LTER PASTA. Used MarcrosExportEML_HTML (working)pie_excel2007_Mar2016.xlsm 3/07/16 3:34 PM for QA/QC to EML 2.1.0
Version 03: August 23, 2018, data and metadata updates to comply with importation to Drupal and LTER PASTA. Used MarcrosExportEML_HTML (working)pie_excel2007_Jan2018.xlsm 6/21/18 3:53 PM for QA/QC to EML 2.1.0
Version 04: August 7, 2019, data and metadata updates to comply with importation to DEIMS7 and LTER Data Portal. Used MarcrosExportEML_HTML (working)pie_excel2007_Jun2019.xlsm 6/7/19 12:58 PM for QA/QC to EML 2.1.0



